miRNAs in human melanoma
High-throughput miRNA profiling of human melanoma blood samples
Petra Leidinger [1,+], Andreas Keller [2,3,+], Anne Borries [2], Jörg Reichrath [4], Knut Rass [4], Sven Uwe Jager [5], Hans-Peter Lenhof [6], Eckart Meese [1,*]
1 Institute of Human Genetics, Medical School, Saarland University, 66421 Homburg/Saar, Germany
2 febit biomed gmbh, Im Neuenheimer Feld 519, 69120 Heidelberg, Germany
3 Biomarker Discovery Center Heidelberg, 69120 Heidelberg, Germany
4 Clinic for Dermatology, Venerology and Allergology, Medical School, Saarland University, 66421 Homburg/Saar, Germany
5 Praxis für Dermatologie, 66280 Sulzbach/Saar, Germany
6 Center for Bioinformatics, Saarland University, Building E 1 1, 66041 Saarbruecken, Germany
+ Contributed equally
* Corresponding author
Abstract
MicroRNA (miRNA) signatures are not only found in cancer tissue but also in blood of cancer patients. Specifically, miRNA detection in blood offers the prospect of a non-invasive analysis tool. Using a microarray based approach we screened almost 900 human miRNAs to detect miRNAs that show deregulated expression in blood cells of melanoma patients. We analyzed 55 blood samples, including 20 samples of healthy individuals, 24 samples of melanoma patients as test set, and 11 samples of melanoma patients as independent validation set. A hypothesis test based approach revealed 51 differentially expressed miRNAs, including 21 miRNAs that were downregulated and 30 miRNAs that were upregulated in blood cells of melanoma patients, as compared to blood cells of healthy controls. The test set and the independent validation set of the melanoma samples showed a high correlation of fold changes (0.81). Applying hierarchical clustering and principal component analysis we found that blood samples of melanoma patients and healthy individuals can be well differentiated from each other based on miRNA expression analysis. Using a subset of 16 significantly deregulated miRNAs, we were able to reach a classification accuracy of 97.4%, a specificity of 95% and a sensitivity of 98.9% by supervised analysis. MiRNA microarray data were validated by qRT-PCR. Our study provides strong evidence for miRNA expression signatures of blood cells as useful biomarkers for melanoma.
Introduction
Recently, microRNAs (miRNAs) have been introduced as new cancer markers in the biomarker landscape and are suggested as targets or future therapy approaches [1,2]. Recent proof-of-principle studies indicate that analysis of miRNA expression in sera and peripheral blood cells is a promising approach for a blood-based diagnosis of cancer and other diseases [3-7]. We recently showed that complex miRNA expression patterns, rather than single miRNAs, can serve as biomarker signatures. Specifically, we were able to separate patients with different human diseases, including lung cancer [8] and Multiple Sclerosis [9] from healthy individuals by blood testing.
In this study, we describe a highly specific miRNA expression profile for melanoma patients. Malignant melanomas represent the most aggressive form of skin cancer. Melanoma accounts for about 4% of skin cancer cases but for as many as 74% of all deaths of skin cancer. The 5-year survival rate is as low as 5% for patients with advanced melanoma [10].
Methods
We used the Geniom Real Time Analyzer microarray platform (febit biomed GmbH, Heidelberg) to analyze all human miRNAs as annotated in the Sanger miRBase version 12.0 [11-13]. In total, we analyzed 35 blood samples of melanoma patients and 20 blood samples of healthy individuals, and validated 13 miRNAs that showed significant deregulation in the microarray experiments using TaqMan® MicroRNA Assays (Applied Biosystems).
Results
We identified 51 differentially expressed miRNAs, including 21 miRNAs downregulated and 30 miRNAs upregulated in blood cells of melanoma patients compared to blood cells of healthy individuals. We validated the microarray results of 13 of the above mentioned 51 miRNAs using qRT-PCR. Analyzing ten samples of melanoma patients and ten samples of healthy controls, we found a correlation of 0.93 between the microarray data and the qRT-PCR data.
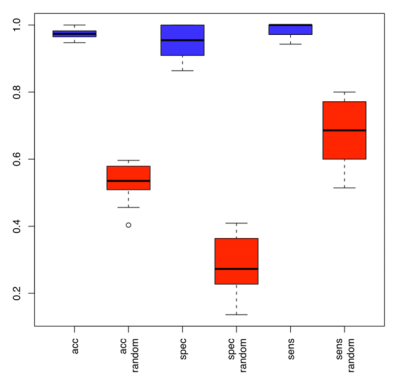
Regarding patient classification, the best accuracy has been obtained using a subset that consisted of 16 miRNAs, including hsa-miR-186, hsa-let-7d*, hsa-miR-18a*, hsa-miR-145, hsa-miR-99a, hsa-miR-664, hsa-miR-501-5p, hsa-miR-378*, hsa-miR-29c*, hsa-miR-1280, hsa-miR-365, hsa-miR-1249, hsa-miR-328, hsa-miR-422a, hsa-miR-30d, and hsa-miR-17*. By using these 16 miRNAs, we separated melanoma from healthy controls with outstanding accuracy, specificity and sensitivity of 97.4%, 95.0% and 98.9%, respectively (Figure 1).
Figure 1: Accuracy, specificity and sensitivity by which melanoma patients are identified based on miRNA profiling of blood. Blue boxes show the classification accuracy, specificity and sensitivity as determined by repeated cross-validation for the subset of 16 miRNAs. Red boxes show the respective accuracy, specificity and sensitivity for the permutation test.
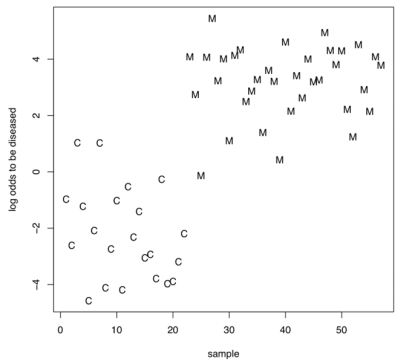
An example of a classification result is shown in Figure 2. In detail, we determined the probability of being a melanoma patient or a healthy control based on the miRNA expression profiles of blood cells. These results further demonstrate that miRNA expression profiling of blood cells can separate melanoma patients from healthy individuals with high sensitivity and specificity.
Figure 2: Example for the classification of melanoma patients and healthy individuals based on the miRNA expression profiling of blood cells for the 16 miRNAs as detected by the subset selection. The logarithm of the quotient of the probability to be a melanoma patient and the probability to be a healthy individual is given on the y-axis for each control (C) and each melanoma (M). If this quotient is greater than one e.g. the logarithm is greater zero the sample is more likely to be a melanoma sample than a control sample.
Discussion
One of the major challenges of improving melanoma treatment is the identification of appropriate markers for the earliest possible detection of the primary lesion. Since melanoma is a very heterogeneous disease, complex biomarker profiles appear to be best suited for the task of early tumor detection and the monitoring of high-risk patients. miRNA expression profiling especially of human blood has the potential to serve as future tumor biomarker test. In this study we provide first evidence for the potential of miRNA expression profiles to distinguish tumor patients from healthy control subjects based on the analysis of peripheral blood cells. Blood-borne miRNA signatures may be ideally suited as prognostic, predictive, or pharmacodynamic biomarkers not only for melanoma but also for other diseases. miRNA expression patterns of cancers or non-cancer diseases might be used to tell these diseases apart form controls and possibly apart from each other.
References
- Cho WC: MicroRNAs in cancer - from research to therapy. Biochim Biophys Acta 2009.
- Cho WC: MicroRNAs: Potential biomarkers for cancer diagnosis, prognosis and targets for therapy. Int J Biochem Cell Biol 2009
- Wang QZ, Xu W, Habib N, Xu R: Potential uses of microRNA in lung cancer diagnosis, prognosis, and therapy. Curr Cancer Drug Targets 2009, 9(4):572-594.
- Wang K, Zhang S, Marzolf B, Troisch P, Brightman A, Hu Z, Hood LE, Galas DJ: Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc Natl Acad Sci U S A 2009, 106(11):4402-4407.
- Cortez MA, Calin GA: MicroRNA identification in plasma and serum: a new tool to diagnose and monitor diseases. Expert Opin Biol Ther 2009, 9(6):703-711.
- Chin LJ, Slack FJ: A truth serum for cancer--microRNAs have major potential as cancer biomarkers. Cell Res 2008, 18(10):983-984.
- Gilad S, Meiri E, Yogev Y, Benjamin S, Lebanony D, Yerushalmi N, Benjamin H, Kushnir M, Cholakh H, Melamed N et al: Serum microRNAs are promising novel biomarkers. PLoS One 2008, 3(9):e3148.
- Keller A, Leidinger P, Borries A, Wendschlag A, Wucherpfennig F, Scheffler M, Huwer H, Lenhof HP, Meese E: miRNAs in lung cancer - studying complex fingerprints in patient's blood cells by microarray experiments. BMC Cancer 2009, 9:353.
- Keller A, Leidinger P, Lange J, Borries A, Schroers H, Scheffler M, Lenhof HP, Ruprecht K, Meese E: Multiple sclerosis: microRNA expression profiles accurately differentiate patients with relapsing-remitting disease from healthy controls. PLoS One 2009, 4(10):e7440.
- Mueller DW, Bosserhoff AK: Role of miRNAs in the progression of malignant melanoma. Br J Cancer 2009, 101(4):551-556.
- Griffiths-Jones S: miRBase: the microRNA sequence database. Methods Mol Biol 2006, 342:129-138.
- Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ: miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 2006, 34(Database issue):D140-144.
- Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ: miRBase: tools for microRNA genomics. Nucleic Acids Res 2008, 36(Database issue):D154-158.